Snakeviz - Visualize Profiling Results in Python¶
The profiling is an important form of analysis which can be used to analyze time or space complexity of code. The Python has many profiling libraries like cProfile, profile, line_profiler, etc to analyze time complexity and memory_profiler, memprof, guppy/hpy, etc to analyze space complexity. The results generated by profiling libraries like cProfile generally log files with many lines each explaining the usage time of various function calls. If the function is very deep and has many lines of code than analyzing such log files can be a very tedious task.
Data visualization is a process where we can represent a lot of data and the human eye can easily catch patterns as well as understand data better. The Python has a library called snakeviz which can take profiling files generated by cProfile and generate visualization out of it.
If you are interested in learning about cProfile then please feel free to visit our tutorial on the same:
The snakeviz has two visualization styles for visualizing profiling results.
- Icicle chart
- Sunburst Chart
The icicle chart uses the width of the rectangle to represent the time taken by a function and the sunburst chart uses an angular extent of arc to represent the time taken by a function. The function which calls other functions will have a special child which represents the time taken by that function outside of other functions which it calls. The only function which calls other functions will have such a child.
We'll now explain the usage of snakeviz through various examples.
Example 1 Snakeviz from Command Line/Shell¶
Our first example which we'll use to explain snakeviz usage is a simple script. It calls main_func() which calls 3 functions and prints results returned by all one by one. All three functions are the same where each generates 100000 random numbers between 1-100 and returns the average of that number. The only difference is the time taken by each function. We have manually introduced a timer that would pause each function for a different amount of time. We'll profile this script using cProfile and then use snakeviz visualize results.
addition.py
import time
import random
def very_slow_random_generator():
time.sleep(5)
arr = [random.randint(1,100) for i in range(100000)]
return sum(arr) / len(arr)
def slow_random_generator():
time.sleep(2)
arr = [random.randint(1,100) for i in range(100000)]
return sum(arr) / len(arr)
def fast_random_generator():
time.sleep(1)
arr = [random.randint(1,100) for i in range(100000)]
return sum(arr) / len(arr)
def main_func():
result = fast_random_generator()
print(result)
result = slow_random_generator()
print(result)
result = very_slow_random_generator()
print(result)
main_func()
We'll use the below command to profile addition.py.
python -m cProfile -o addition.prof addition.py
We can then call the below command to launch snakeviz visualization in the browser.
snakeviz addition.prof
After execution of the above command snakeviz will launch visualization a browser as shown below.
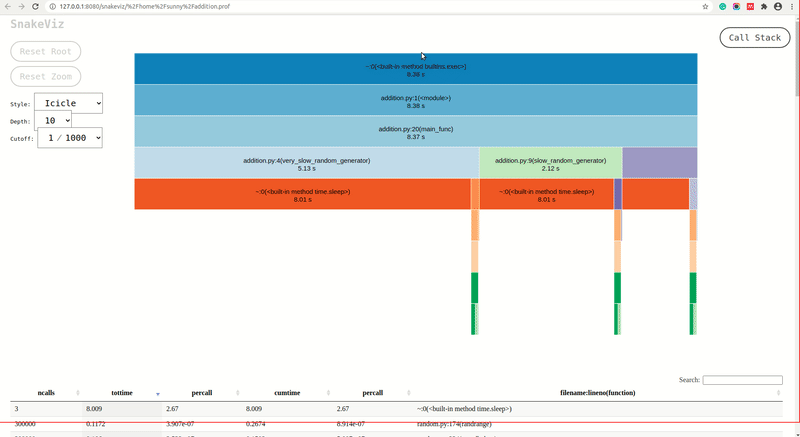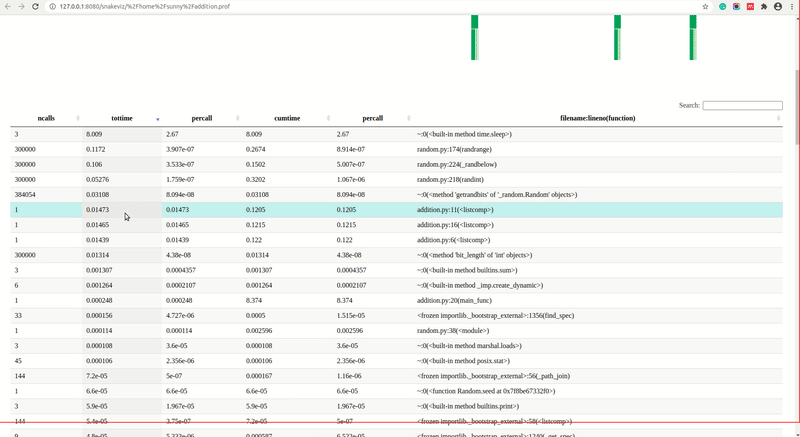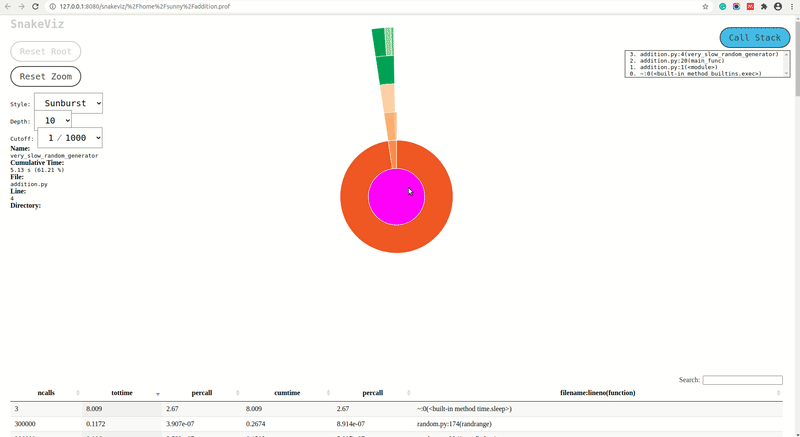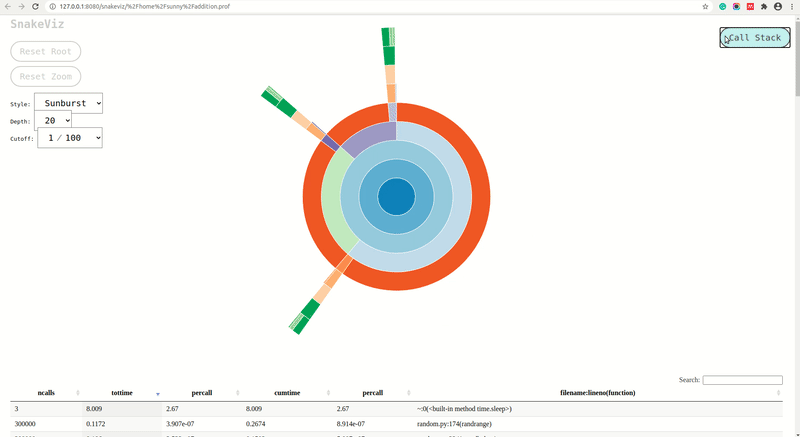
As we can see from the above screencast that snakeviz generated two charts by default and we can change the chart from the dropdown. All individual lines of profiling will be shown as the table below visualization.
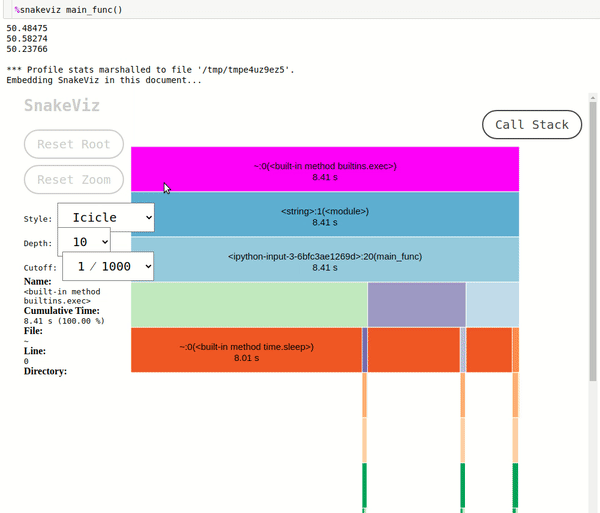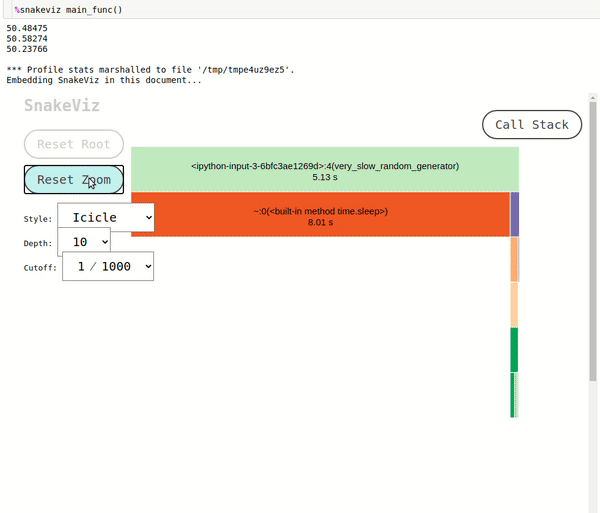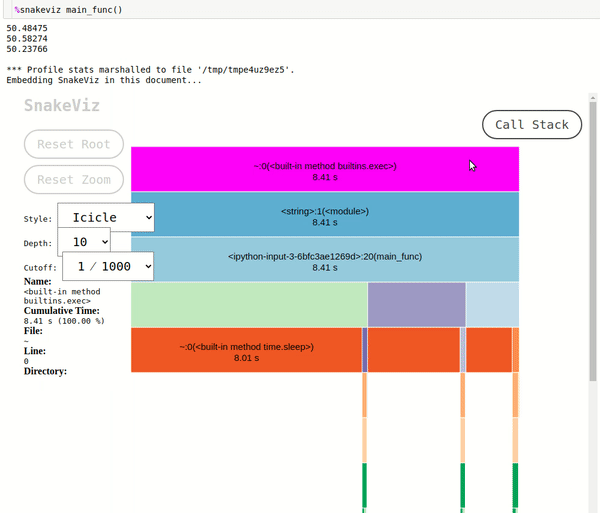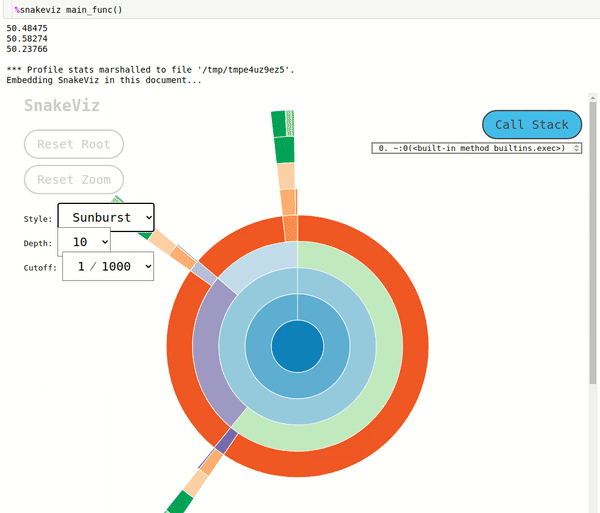
- Icicle Chart: The icicle chart uses rectangles to represent the time taken by functions. We can see that the total time taken by our script is ~8.38 seconds. The rectangles for
fast_random_generator,slow_random_generatorandvery_slow_random_generatorare shown inside of rectangle ofmain_func(). We can further click on a rectangle of any of that functions and it'll be set as root rectangle and all sub-functions called inside it'll be as rectangles below it. The rectangles are organized like a tree structure in a fashion in which they were called and the width of the rectangle represents the time taken by that function in the caller function of that function. - Sunburst Chart: The sunburst chart uses the angular extent of arc to represent the time taken by functions. We can see exactly same way as icicle chart that arc of
fast_random_generator,slow_random_generatorandvery_slow_random_generatorare shown inside extent of arc ofmain_func(). We can click on any arc and that arc will become a root arc and functions called inside it'll be shown as arcs around it.
We'll now explain various components of visualization which can help better understand the visualization.
- Function Information: We can hover over the rectangle in the icicle chart or arc in the sunburst chart and it'll show which information of function which it represents on the left side. It'll show the name of the function, cumulative time, file in which the function is present, line number on which it gets called, and directory of file.
- Call Stack: The
Call Stackbutton present on top right represents the call stack. If we click on any rectangle or arc then the chart will make that node as the root node and all sub-nodes will show functions called inside it. It'll take us deep into that function. If we have gone deep into functions analyzing them one by one and want to know our stack path then we can click on theCall Stackbutton and it'll show us call stack. We can also click on any entry in it and it'll take us to that level. - Stats Table: The stats table looks exactly the same as that of output generated by
cProfile. It'll be displayed below chart. Each of the lines of the state table is clickable representing function calls and when we click on any line that line will be made root node of the chart and all functions called inside it will be shown as sub-nodes. Please feel free to visit our tutorial oncProfilewhich explains what each column means in this table in-depth. - Reset Root: If we have changed the root of the chart by clicking on the row of the stats table than we can click on this button to reset the root to the original.
- Reset Zoom: If we have zoomed into profiling by clicking in on any rectangle or arc then we can reset it by clicking this button.
- Style: The style has two options for two chart types
- Icicle
- Sunburst
- Depth: The
Depthdropdown lets us select how deep into call stack snakeviz goes when creating a visualization. The calls below this depth won't be shown into the chart until we click on one of the top functions which call it. We can increase profile depth and that many levels will be shown in the chart at once. - Cutoff: The
Cutoffdropdown can let us decide whether to display function calls which takes up very little time of their parent’s cumulative time. If we setCutOffto some value then for each function we check for the ratio of its cumulative time and its parent’s cumulative time. If the ratio is less than the cutoff value then that function will be displayed but the chart won't be constructed for sub-functions inside of that function which means that we can't click on that function anymore to see time spent on sub-functions inside it. If we set a high cutoff then the chart will be generated faster as many sub-charts won't be generated.
Example 2: Snakeviz inside Jupyter Notebook¶
We can easily use snakeviz inside of the jupyter notebook as well. We first need to load snakeviz as an extension inside of the notebook and then we can call snakeviz as a line or cell magic command. Below we have loaded snakeviz as an extension.
%load_ext snakeviz
Below we have used the same code as our first example, but when we are calling main_func(), we have used the %snakeviz line command to generate snakeviz visualization for it. It'll generate visualization inside of the notebook.
import time
import random
def very_slow_random_generator():
time.sleep(5)
arr = [random.randint(1,100) for i in range(100000)]
return sum(arr) / len(arr)
def slow_random_generator():
time.sleep(2)
arr = [random.randint(1,100) for i in range(100000)]
return sum(arr) / len(arr)
def fast_random_generator():
time.sleep(1)
arr = [random.randint(1,100) for i in range(100000)]
return sum(arr) / len(arr)
def main_func():
result = fast_random_generator()
print(result)
result = slow_random_generator()
print(result)
result = very_slow_random_generator()
print(result)
%snakeviz main_func()
Below we have generated snakeviz visualization again using the same code but this time using the %%snakeviz cell magic command to explain the usage of it.
%%snakeviz
import time
import random
def very_slow_random_generator():
time.sleep(5)
arr = [random.randint(1,100) for i in range(100000)]
return sum(arr) / len(arr)
def slow_random_generator():
time.sleep(2)
arr = [random.randint(1,100) for i in range(100000)]
return sum(arr) / len(arr)
def fast_random_generator():
time.sleep(1)
arr = [random.randint(1,100) for i in range(100000)]
return sum(arr) / len(arr)
def main_func():
result = fast_random_generator()
print(result)
result = slow_random_generator()
print(result)
result = very_slow_random_generator()
print(result)
main_func()

This ends our small tutorial explaining usage of snakeviz. Please feel free to let us know your views in the comments section.
References¶
- How to Profile Python Code using cProfile & profile?
- Snakeviz Docs
- How to Profile Memory Usage in Python using memory_profiler
- line_profiler: Line by Line Profiling of Python Code
- py-spy - Sampling Profiler for Python Code
- pyinstrument - Statistical Profiler for Python Code
- guppy/heapy - Profile Memory Usage in Python
- yappi - Yet Another Python Profiler
- pympler - Monitor Memory Usage by Python Objects
- pprofile - Deterministic and Statistical Profiler for Python Code
- Scalene - CPU and Memory Profiler for Python Code
- tracemalloc - How to trace memory usage in Python code
 Sunny Solanki
Sunny Solanki
![YouTube Subscribe]() Comfortable Learning through Video Tutorials?
Comfortable Learning through Video Tutorials?
If you are more comfortable learning through video tutorials then we would recommend that you subscribe to our YouTube channel.
![Need Help]() Stuck Somewhere? Need Help with Coding? Have Doubts About the Topic/Code?
Stuck Somewhere? Need Help with Coding? Have Doubts About the Topic/Code?
When going through coding examples, it's quite common to have doubts and errors.
If you have doubts about some code examples or are stuck somewhere when trying our code, send us an email at coderzcolumn07@gmail.com. We'll help you or point you in the direction where you can find a solution to your problem.
You can even send us a mail if you are trying something new and need guidance regarding coding. We'll try to respond as soon as possible.
![Share Views]() Want to Share Your Views? Have Any Suggestions?
Want to Share Your Views? Have Any Suggestions?
If you want to
- provide some suggestions on topic
- share your views
- include some details in tutorial
- suggest some new topics on which we should create tutorials/blogs



 Comfortable Learning through Video Tutorials?
Comfortable Learning through Video Tutorials? Stuck Somewhere? Need Help with Coding? Have Doubts About the Topic/Code?
Stuck Somewhere? Need Help with Coding? Have Doubts About the Topic/Code? Want to Share Your Views? Have Any Suggestions?
Want to Share Your Views? Have Any Suggestions? profiling, snakeviz
profiling, snakeviz